This is a demo store. No orders will be fulfilled.
Epigenetic Modifiers
Introduction to Epigenetics
Epigenetics refers to modifications in gene activity that occur without altering the underlying DNA sequence. Rather than changing the genetic code, these processes regulate whether genes are switched on or off through mechanisms such as chromatin remodeling. This often involves chemical changes to histone proteins or the attachment of methyl or alkyl groups to DNA bases.
Alterations like histone acetylation or methylation can significantly influence chromatin structure, thereby affecting the likelihood of transcription. In a similar way, excessive DNA methylation within a gene’s promoter region can block transcription and prevent gene expression. Because of this, enzymes that control these modifications—such as histone deacetylases and DNA methyltransferases—are being investigated as therapeutic targets, particularly for their potential in treating various forms of cancer.

DNA Methyltransferase (DNMTs)

DNA methyltransferases (DNMTs) primarily catalyze the addition of methyl groups to CpG sites within DNA. In mammals, three enzymatically active DNMTs have been identified: DNMT1, DNMT3A, and DNMT3B. Methylation at promoter regions generally silences gene expression by blocking the ability of transcription factors to bind to DNA. Beyond this, methylated DNA can be recognized by methyl-CpG–binding domain proteins, which in turn recruit histone-modifying enzymes. These enzymes compact chromatin, providing an additional route for gene repression.
In certain cancers, such mechanisms contribute to reduced activity of tumor suppressor genes, promoting uncontrolled cellular proliferation. To counteract this, several agents that inhibit DNMT activity have been developed, including RG-108, mithramycin, and azacytidine.
| Aladdin Cat # | Product Name | Description | Purity |
A119533 | 5-Aza-2’-deoxycytidine (Decitabine) | Inhibits DNMT1/3A/3B | ≥98% |
A755608 A100625 A408358 | Azacitidine | Inhibits DNMT1/3A/3B | BioReagent,≥98%, 10mM in DMSO |
B117979 B131602 | Bisdemethoxycurcumin | Inhibits DNMT1 | ≥98% , Analytical standards |
C109404 C109403 C109402 C423343 | Chlorogenic Acid | Inhibits DNMT | ≥98%, Analytical standards≥98%,≥95%, 10mM in DMSO |
E107403 E107404 E409009 | Epigallocatechin Gallate | Inhibits DNMT1 | Analytical standards, ≥98%, 10mM in DMSO |
F107712 F107713 F156691 F409163 | Fisetin | Inhibits DNMT1 | Analytical standards, ≥98%, ≥98%, ≥96%, 10mM in DMSO |
Lomeguatrib | Inhibits O6-methylguanine-DNMT (MGMT) | ≥98%, 10mM in DMSO | |
Mithramycin/ Plicamycin | Inhibits DNMT1 | ≥95% | |
O6-Benzylguanine | Inhibits O6-methylguanine-DNMT (MGMT) | ≥98%, 10mM in DMSO | |
RG-108 | Inhibits DNMT | ≥98%, 10mM in DMSO | |
Sinefungin | Inhibits DNMT | ≥95% | |
Sorafenib | Inhibits DNMT activity | ≥99%, 10mM in DMSO | |
Theaflavin-3,3’-digallate | Inhibits DNMT | ≥98% |
Histone Methyltransferase Inhibitors
Histone methyltransferases (HMTs) catalyze the addition of methyl groups to lysine and arginine residues on histone proteins, most notably histones H3 and H4. This methylation alters the histone’s chemical properties, reducing its positive charge and slightly loosening its interaction with DNA. As a result, the DNA becomes more accessible, which can promote transcriptional activity. In this way, HMTs may enhance gene expression by facilitating transcription of DNA sequences associated with less compact chromatin.
However, histone methylation is context-dependent and can also repress transcription. Depending on the specific histone residue involved, methylation may obstruct transcription factor binding sites or encourage chromatin condensation, thereby silencing genes. In certain cancers, aberrant activity of methyltransferases such as EZH2 and DOT1L contributes to the repression of tumor suppressor genes. Targeting these enzymes with inhibitors—including EPZ5676, EPZ005687, and GSK126—has demonstrated anticancer efficacy in multiple in vitro and in vivo studies.


Aladdin Cat # | Product Name | Description | Purity |
EPZ005687 | Inhibits EZH2 | ≥98%, 2mM in DMSO | |
EPZ5676 | Inhibits DOT1L | ≥98%, 10mM in DMSO | |
EPZ6438 | Inhibits EZH2 | ≥99%, 2mM in DMSO | |
GSK126 | Inhibits EZH2 | ≥98%, 2mM in DMSO | |
GSK343 | Inhibits EZH2 | ≥98%, 2mM in DMSO | |
Inhibits EZH2 | ≥99%, 10mM in DMSO |
Histone Deacetylases (HDACs)
|
Aladdin Cat #
|
Product Name
|
Description
|
Purity
|
AK-7 | Inhibits SIRT2 (HDAC class III), brain penetrant | ≥98%, 10mM in DMSO | |
Apicidin | Inhibits HDAC (broad spectrum, class I/II) | ≥98% | |
Belinostat | Inhibits HDAC | ≥98%, 10mM in DMSO | |
Butyric Acid Sodium | Inhibits HDAC | 10mM in Water, BioReagent ≥99%,≥98% | |
Cambinol | Inhibits SIRT1 (HDAC class III) | ≥98%, 10mM in DMSO | |
Curcumin | Decreases expression of HDAC3 (class I) | Natural extraction(isomer mixture), 0.1% in 95% Ethanol, from Synthetic ≥98%, analytical standard, ≥65%,≥95%,≥75%(Curcumin) total curcumin content, from Curcuma longa (Turmeric),powder, 10mM in DMSO | |
Entinostat | Inhibits HDAC1 (class I) | ≥98%, 10mM in DMSO | |
Isoliquiritigenin | Inhibits HDAC (class I/IIA) | ≥98%, analytical standard | |
LBH-589 | Inhibits HDAC1/2/3/11 (class I) | ≥98%, 10mM in DMSO | |
MGCD-0103 | Inhibits HDAC | ≥98%, 10mM in DMSO | |
Mycophenolic Acid | Inhibits HDAC | ≥98%, 10mM in DMSO | |
Phenylbutyrate | Inhibits HDAC | ≥98% | |
Romidepsin | Inhibits HDAC | ≥98%, 10mM in DMSO | |
Salermide | Inhibits SIRT1/2 (HDAC class III) | ≥98%, 10mM in DMSO | |
Sirtinol | Inhibits SIRT1/2 (HDAC class III) | ≥98%, 10mM in DMSO | |
Scriptaid | Inhibits HDAC (broad spectrum) | ≥99%, 10mM in DMSO | |
Sorafenib | Decreases expression of HDAC1/2/4/5/8 (classI/IIA) | ≥99% | |
TMP-269 | Inhibits HDAC (class II) | ≥98%, 10mM in DMSO | |
Tozasertib | Decreases expression of HDAC | ≥98%, 10mM in DMSO | |
Trichostatin A | Inhibits HDAC1/3/4/6/10 (class I/IIA/IIB) | ≥98%, ≥95% | |
Tubacin | Inhibits HDAC6/10 (class IIB) | ≥96% | |
Tubastatin A HCl | Inhibits HDAC6/10 (class IIb) | ≥98%, 10mM in DMSO | |
n-Valeric Acid | Inhibits HDAC | ≥99%, analytical standard, CP ≥98%, Standard for GC ≥99.5%(GC) | |
Valproic Acid Na+ Salt | Inhibits HDAC1 (class I) | ≥98%, 10mM in DMSO | |
Vorinostat (SAHA) | Inhibits HDAC1/2/3/6 (class I/IIB) | ≥99%, 10mM in DMSO |
Histone deacetylases (HDACs) function by removing acetyl groups from N-acetyl lysine residues on histone proteins. This deacetylation increases the positive charge on histones, strengthening their interaction with the negatively charged DNA backbone. The result is a more compact chromatin structure, which reduces the likelihood of transcription. Through this mechanism, HDACs can suppress the expression of genes involved in apoptosis and tumor suppression.
HDACs are categorized into four major classes according to their cellular distribution and biological roles. Class I HDACs (isoforms 1, 2, 3, and 8) are predominantly nuclear, whereas Class II HDACs (isoforms 4, 5, 6, 7, 9, and 10) shuttle between the nucleus and cytoplasm. Inhibition of HDAC activity has shown therapeutic potential, particularly in oncology, where HDAC inhibitors enhance the effects of other chemotherapeutic agents. This strategy has been especially effective in treating leukemias and lymphomas. Examples of HDAC inhibitors include vorinostat, trichostatin A, scriptaid, and phenylbutyrate.
Aurora Kina
|
Aladdin Cat #
| Name |
Description
| Purity |
(Barasertib) | Determined to be the most selective available AurB inhibitor in a 2015 study. | ≥97% | |
CYC-116 | Inhibits AurA and AurB. Induces apoptosis in multiple myeloma cells in combination with matrine. | ≥98%, 10mM in DMSO | |
GSK-1070916 | Inhibits AurB and AurC. | ≥98%, 10mM in DMSO | |
MLN8237 (Alisertib) | Selective AurA inhibitor. Eff ective in treating models of neruoblastoma, acute lymphoblastic leukemia, and sarcoma. | ≥98%, 10mM in DMSO | |
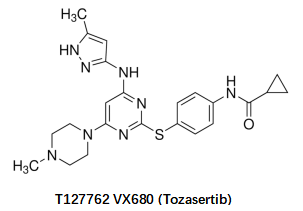
VX680 (Tozasertib) | AurA inhibitor with some AurB inhibitory eff ect. | ≥98%, 10mM in DMSO | |
ZM-447439 Trihydrate | Inhibits AurA and AurB. Limits migration of MCF-7 human breast cancer cells. | ≥98%, 10mM in DMSO |
Aurora kinases comprise a family of serine/threonine kinases that play essential roles in regulating mitosis. Three members—Aurora A, Aurora B, and Aurora C—carry out distinct but complementary functions in chromatid segregation and other aspects of cell division. Overexpression of these kinases has been observed in numerous cancers, highlighting their potential as therapeutic targets.
Inhibition of Aurora kinases triggers apoptosis through specific mechanisms unique to each isoform. Blocking Aurora A disrupts mitotic spindle formation, while inhibition of Aurora B interferes with proper chromosome alignment. The role of Aurora C is less well defined, as it is normally active in meiotic cells; however, emerging evidence suggests it also exhibits oncogenic potential. To date, most drug development efforts have focused on small-molecule inhibitors directed against Aurora A and Aurora B.


Aladdin: https://www.aladdinsci.com/
 首页
首页 400-620-6333
400-620-6333
